Could Dogs Hold the Key to Understanding Human Diabetes? My Genetic Quest!
What if studying dogs could help us better understand — and treat — metabolic disorders in humans? That's the question driving my latest research project, in which I designed primers to analyze the canine version of the GLP1R gene, searching for genetic variants that mirror those linked to diabetes and metabolic disease in people.
Why GLP-1 Matters
To understand my research, it helps to know a little about GLP-1 — glucagon-like peptide-1. This hormone is produced by specialized cells in your gut called L-cells every time you eat. Think of GLP-1 as a master metabolic coordinator: it signals the pancreas to release insulin while simultaneously tapping the brakes on glucagon release. The liver then responds to those signals by carefully adjusting how much glucose it stores or releases into the bloodstream. The result is a finely tuned system that keeps blood sugar levels in a healthy range.
Disruptions in this system — particularly in the gene encoding the GLP-1 receptor (GLP1R) — have been linked to metabolic disorders in humans. Last year, I researched specific variants in the human GLP1R gene associated with these conditions. This year, I turned my attention to dogs.
The Research Question Nobody Had Asked
Here's the surprising part: despite dogs being one of the most medically studied non-human species, nobody had thoroughly examined whether dogs carry the same GLP1R variants that are associated with metabolic disease in humans. My goal was to design the primers and protocols that would make answering that question possible.
The four human SNP sites I focused on — rs6923761, rs10305420, rs10305492, and rs2268641 — are established markers in the scientific literature. My challenge was to find the corresponding locations in the canine genome and build the tools to sequence them.
From Shelter Dogs to DNA Sequences: The Methods
My research journey began, fittingly, with dogs, specifically, canine samples obtained through a relationship my mentor at Cold Spring Harbor Laboratory had established with Brookhaven Animal Shelter.
DNA Extraction came first. I used Chelex extraction, the optimal method for the types of samples I had, to prepare PCR-ready DNA from each animal.
Primer Design was the hardest step of the project. Using GenBank's Comparative Genome Viewer, I aligned the human and dog GLP1R genes to pinpoint where the human SNP sites corresponded in the canine genome. I then confirmed via BLAST that the GLP1R sequence is conserved across dog breeds — good news, because it means the primers I designed should work regardless of dog breeds. Finally, I used Primer-BLAST to design and optimize primers targeting approximately 200-base-pair windows around each of the four sites.
PCR Amplification followed a precise thermocycling protocol:
Denaturation at 95°C,
Annealing at 55°C, and
Elongation at 72°C — each step 30 seconds.
Over 35 cycles, this process produced millions to billions of copies of the four target regions.
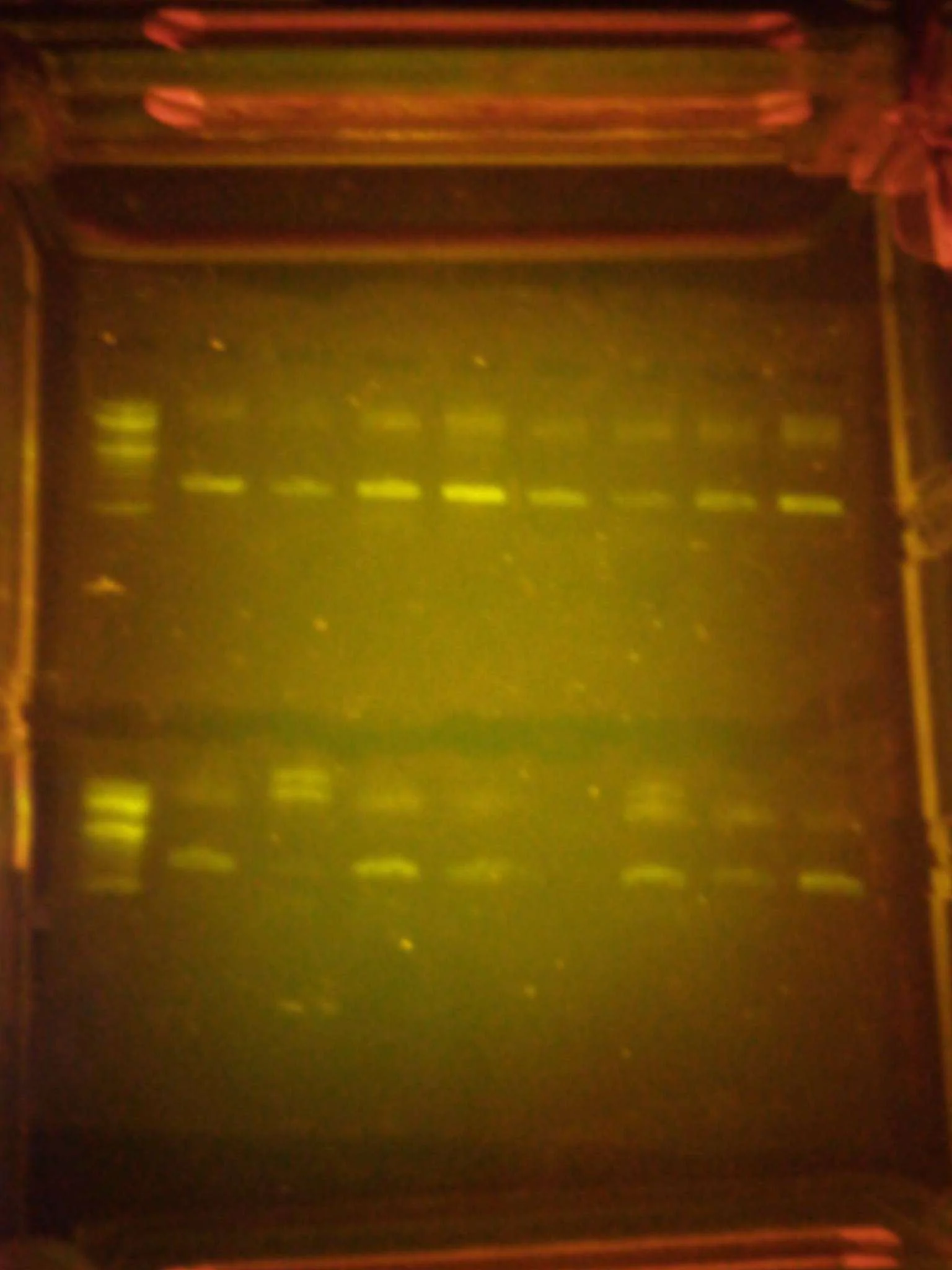
Gel Electrophoresis:
Let me verify the PCR worked, by checking whether DNA fragments migrated to the expected positions on the gel relative to a DNA ladder. Samples that showed clear bands moved on to the next step.
Sanger Sequencing which was performed by an external lab revealed the exact nucleotide sequence in each sample. After that I used DNA Subway to trim low-quality bases, review chromatograms, pair forward and reverse reads into consensus sequences, and then BLAST and MUSCLE-align those sequences to confirm they matched the canine GLP1R gene and targeted the regions I intended.
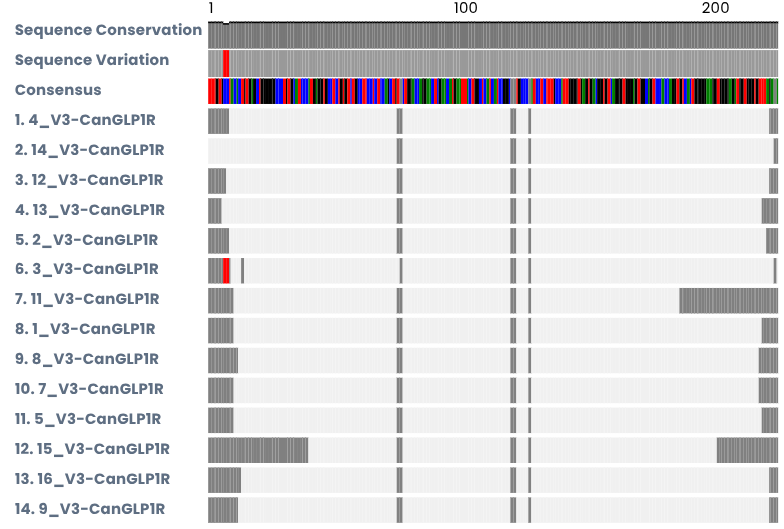
Muscle Alignment
This is an image of an muscle alignment that shows sequence quality
What I Found
The results were a mix of technical success and scientifically interesting silence.
Three out of four primer sets successfully amplified their target regions. One set requires primer optimization before it can be used reliably. One sample — Sample #10 — failed across all primer sets and was excluded due to DNA quality issues.
When I examined the sequenced regions, I found that nearly all samples were genetically identical at these positions: no SNPs were detected. But this isn't a negative result in the traditional sense. My samples were small and randomly selected. I wasn't expecting to find a clear disease association. The primary goal was always to validate that these regions could be amplified and sequenced in dogs, and on that front, the work succeeded.
What Comes Next
The real science starts now. With validated primers and protocols in hand, the next step is to apply them to a larger, more targeted population of dogs that have been diagnosed with diabetes alongside healthy controls. That comparison is what will reveal whether these GLP1R variants exist in dogs at all, and if so, whether they track with disease.
The implications could ripple in two directions. For veterinary medicine, identifying genetic risk factors for canine diabetes could transform how we screen for and manage the condition. For human medicine, dogs that naturally share our GLP1R variants could become powerful models for testing individualized therapies including the GLP-1 receptor agonists that have recently transformed diabetes and obesity treatment in humans.
Science as Problem-Solving
This project taught me things no textbook could. Sourcing approved research facilities, mastering complex lab techniques, navigating bioinformatics tools, every obstacle required adapting the plan, leaning on mentors, and discovering what real-world science actually looks like. It's messier, slower, and more collaborative than it appears from the outside.
But it's also more meaningful. The question of whether dogs and humans share the same metabolic vulnerabilities is more than academic. If the answer is yes, it could change how we treat two species at once.